Local Access
For New R Users:
Download R and RStudio (Free).
Open RStudio. It will look like this:
Click to Expand
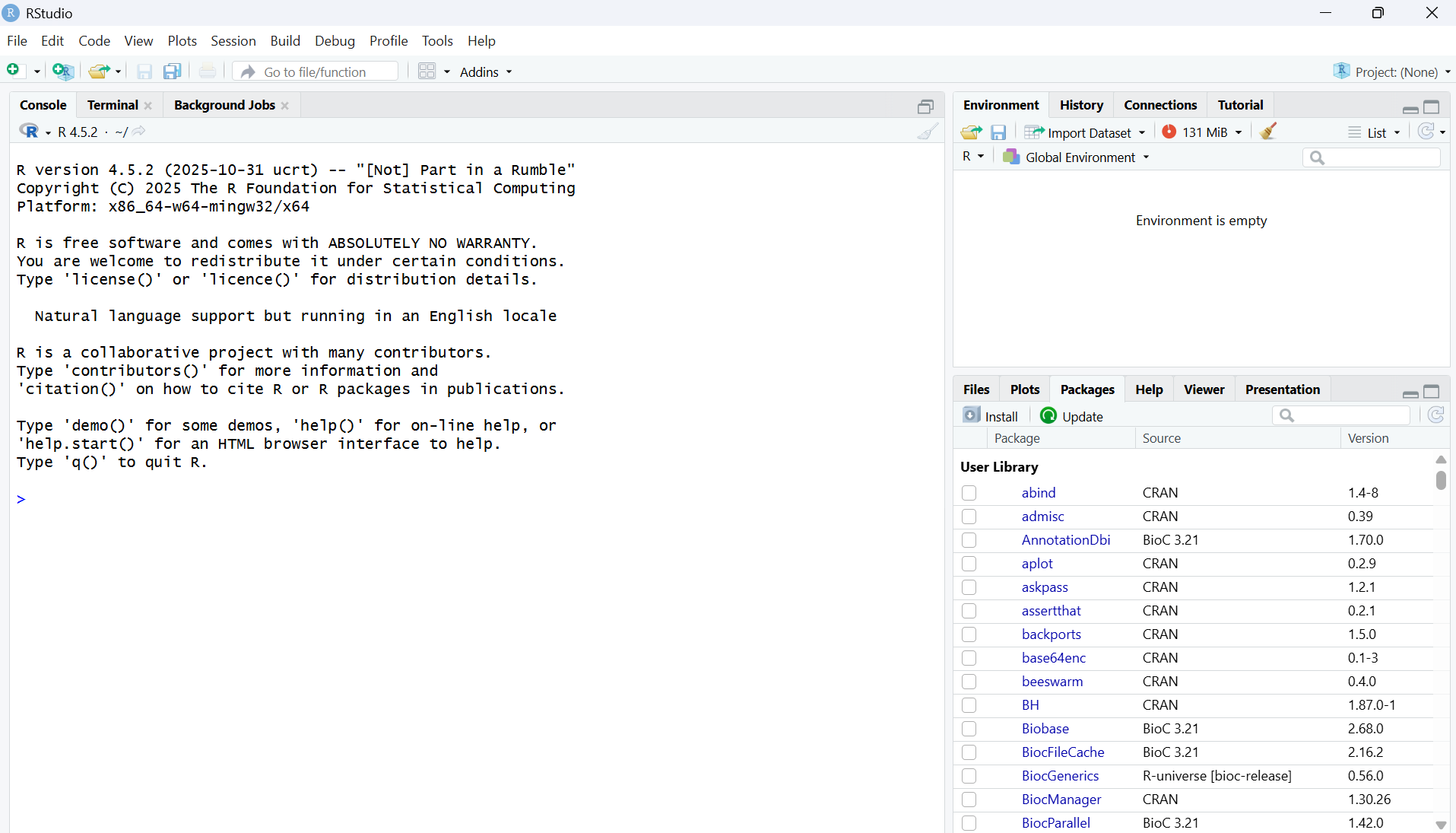
- Within the Console, copy and paste these pieces of code, then press Enter. The console is the panel on the left and starts with >
install.packages('FRDATranscriptomicAtlas', repos = c('https://marniemaddock.r-universe.dev', 'https://cloud.r-project.org'))Once the installation has completed, a > will appear in the console. It will say something similar to ’the downloaded binary packages are in …`. You may recieve a warning saying that Rtools is required to build packages. This is not an issue and can be ignored.
Click to Expand
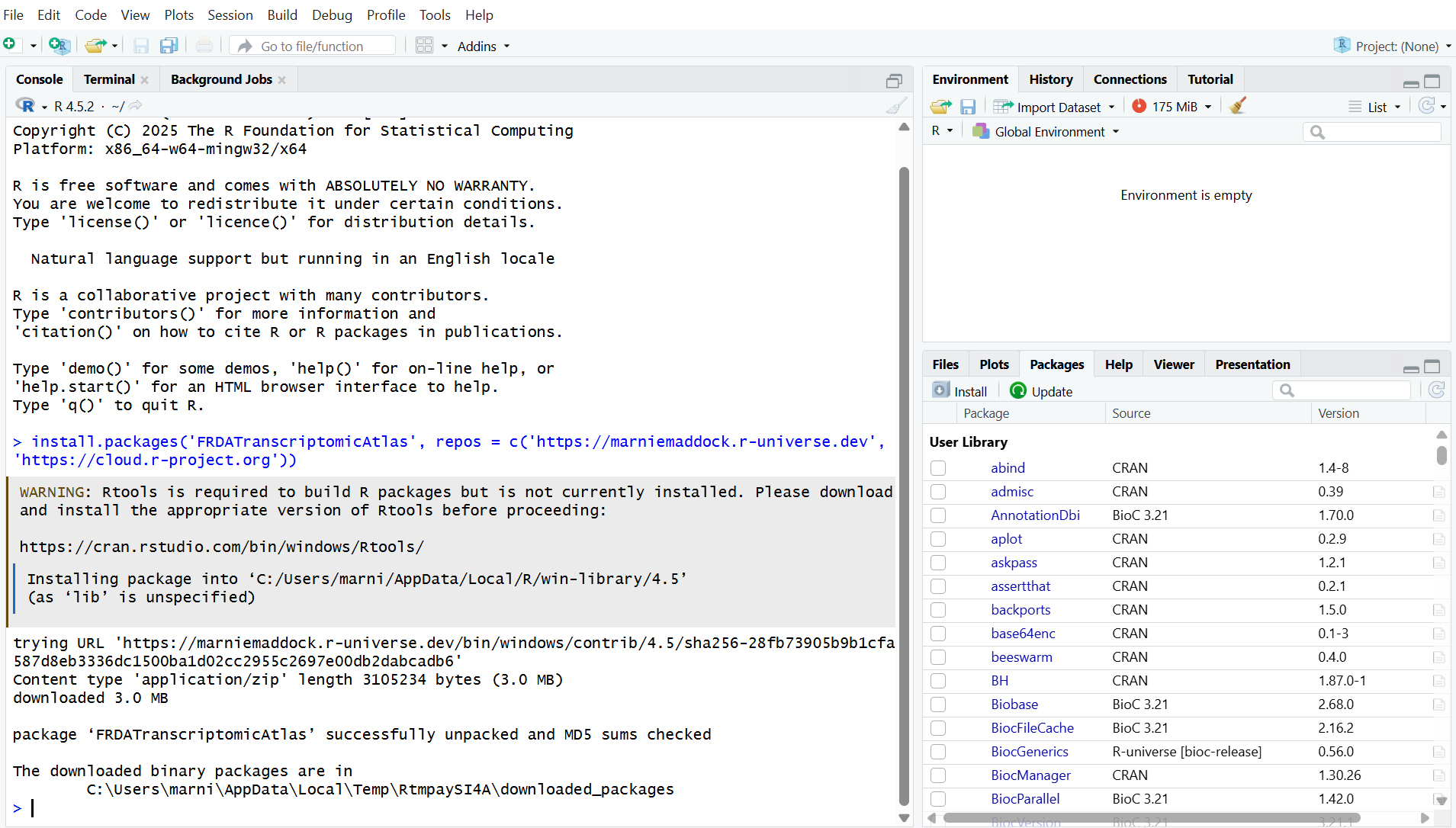
Now run:
FRDATranscriptomicAtlas::install_deps()This will download additional packages required to render heatmaps in the app. Once that has been completed, a > will appear in the console again.
FRDA Transcriptomic Atlas has been downloaded, but it has not been loaded into the R session. To do this, type this line of code in the Console and press control/command enter to run it:
library(FRDATranscriptomicAtlas)This itself will not open the application. To open the application, run:
run_app()This should open the FRDA Transcriptomic Atlas app in a new window. If not, the http address given in your console following “Listening on” can be copied into a web browser to open FRDA Transcriptomic Atlas.
Click to Expand
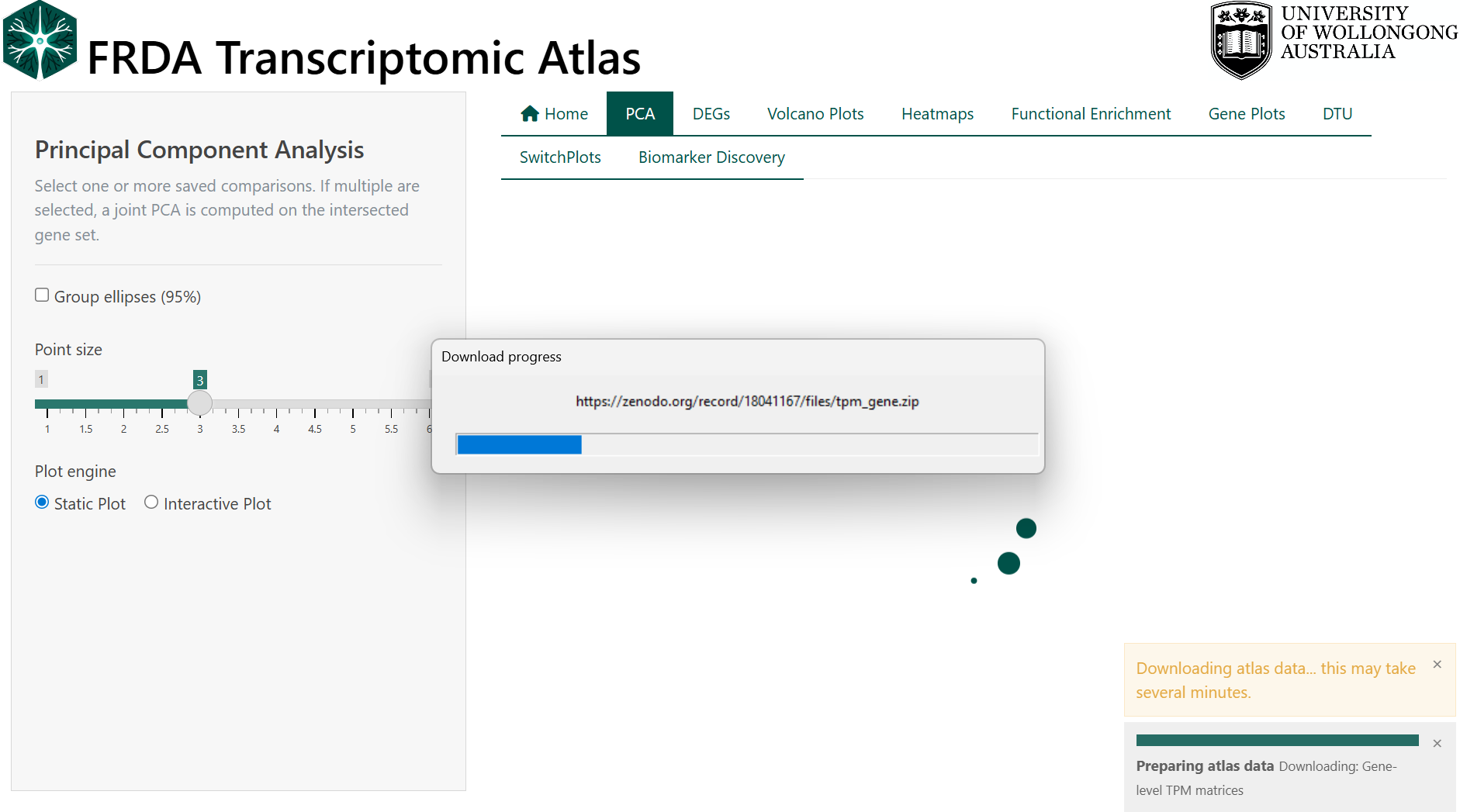
The atlas will then proceed to download the necessary data files. This may take several minutes depending on your internet speed. A progress bar will be displayed. This will only occur the first time you run the app. Once completed, the message `` will appear.
Click to Expand
Once Installed:
Once the FRDA Transcriptomic Atlas app has been installed on the computer, they do not need to be redownloaded unless there are updates i.e. you do not need to run install.packages('FRDATranscriptomicAtlas', repos = c('https://marniemaddock.r-universe.dev', 'https://cloud.r-project.org')) again.
To run FRDA Transcriptomic Atlas in a new R session use (i.e. if you close R and open it again):
library(FRDATranscriptomicAtlas)
run_app()The app will open in a new window. In the top corner you can click open in browser button. This will open the app in your default web browser. You can control the zoom settings of the app in this window.
For Existing R Users:
- Open RStudio and Run:
install.packages('FRDATranscriptomicAtlas', repos = c('https://marniemaddock.r-universe.dev', 'https://cloud.r-project.org'))
FRDATranscriptomicAtlas::install_deps()
library(FRDATranscriptomicAtlas)
run_app()- FRDA Transcriptomic Atlas will open in a new window.